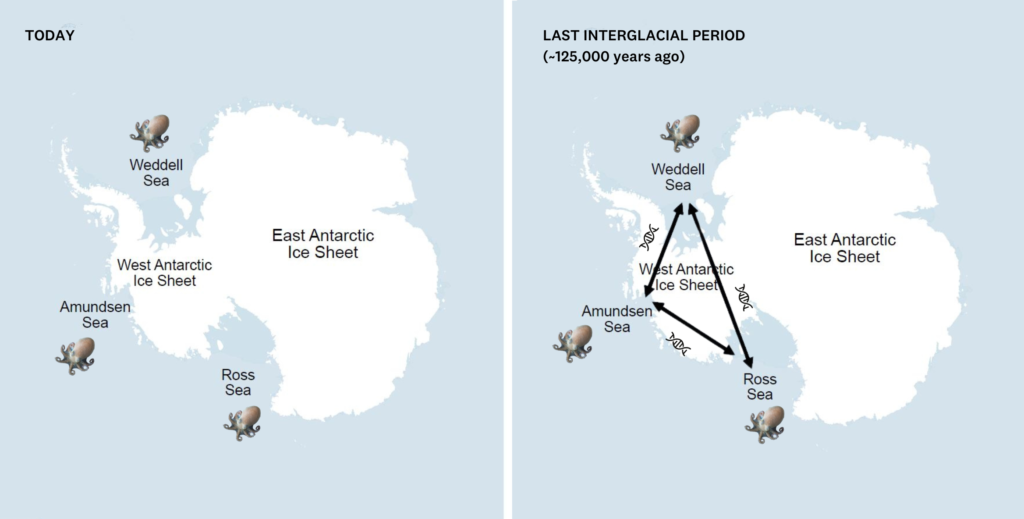
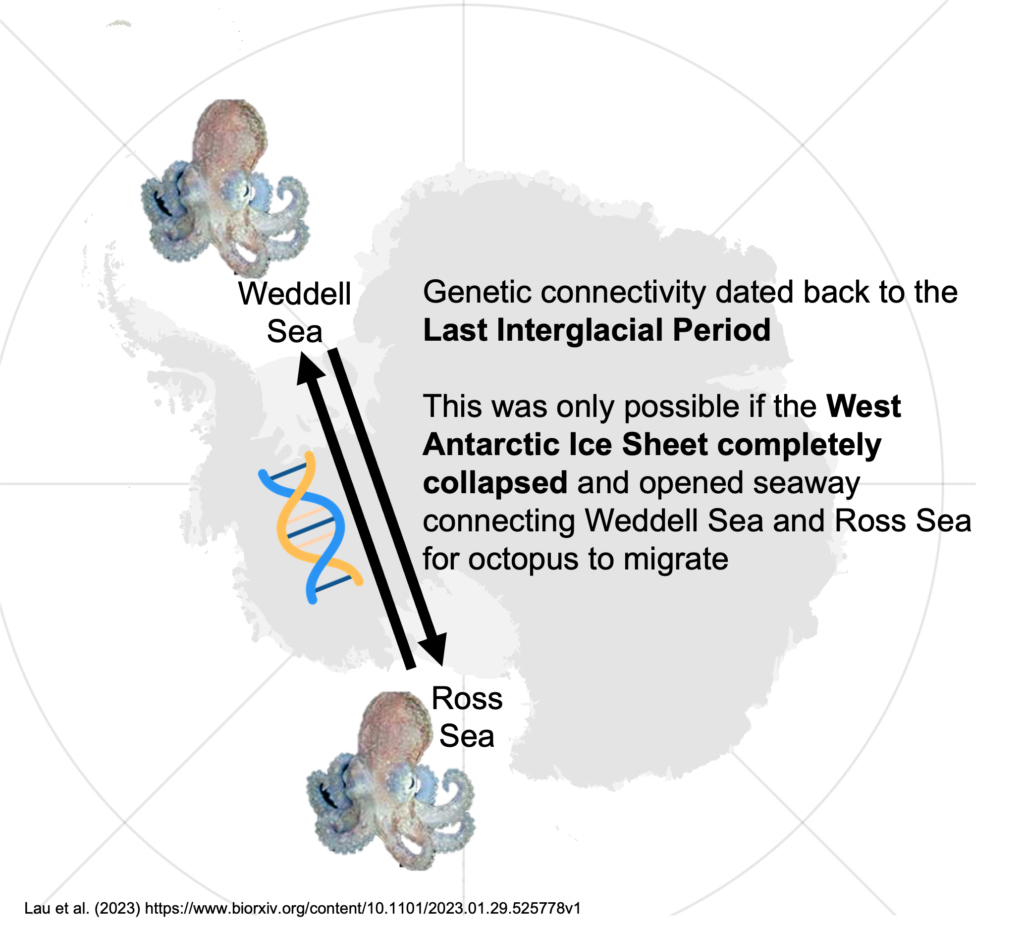
New research published in Current Biology, led by Sally Lau, shows that maintaining and adopting proposed marine protected areas (MPAs) in the Southern Ocean could almost double the protection of genetic hotspots from 28% to ~54%.
These hotspots are home to species with high genetic diversity, which means they are more resilient and better able to adapt to climate and environmental change.
The work provides a new layer of evidence to support the work of CCAMLR in identifying and establishing MPAs.
To conduct the study the team synthesised all the publicly available genetic data from Southern Ocean seafloor species collected over decades through major international initiatives, such as the International Polar Year.
It highlights the value of sustained investment in internationally collaborative Antarctic research. The research was supported by Securing Antarctica’s Environmental Future.
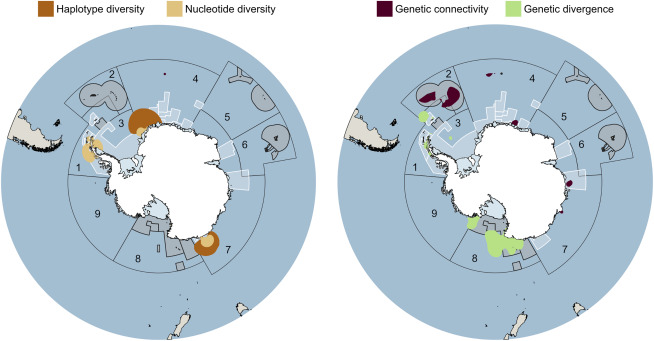
Hotspots of haplotype diversity, nucleotide diversity, genetic divergence, and genetic similarity across the Southern Ocean, based on population-COI data averaged across nine marine invertebrate higher taxa comprising 28 species. Each panel contains a large map reflecting the distribution of hotspots in relation to established (gray shaded areas) and proposed (white shaded areas) MPAs within CCAMLR MPA planning domains 1–9.